Abstract
This paper introduces the Neuroplastic Self-Organizing Map (NPSOM), an unsupervised learning approach that enhances the traditional Self-Organizing Map (SOM) with neuroplasticity features. NPSOM incorporates mechanisms for synaptic decay, neurogenesis, and memory forgetting to improve the adaptability and performance of the SOM. By mimicking the brain’s ability to strengthen certain neurons’ synaptic connections and forget unnecessary information, NPSOM adapts to changing data distributions while retaining important information. The experimental results show that NPSOM outperforms MOSEGCN (best baseline) by 0.3% on the BRCA dataset while delivering similar performance on the ROSMAP dataset in terms of classification accuracy.
Methodology: Neuroplasticity in SOMs
NPSOM builds upon the traditional SOM by introducing three key biological-inspired mechanisms:
- Synaptic Decay: Mimics the weakening of unused neural connections over time.
- Neurogenesis: Allows the model to create new neurons to adapt to new data patterns.
- Memory Forgetting: Enables the model to discard irrelevant or redundant information, focusing on the most salient features.
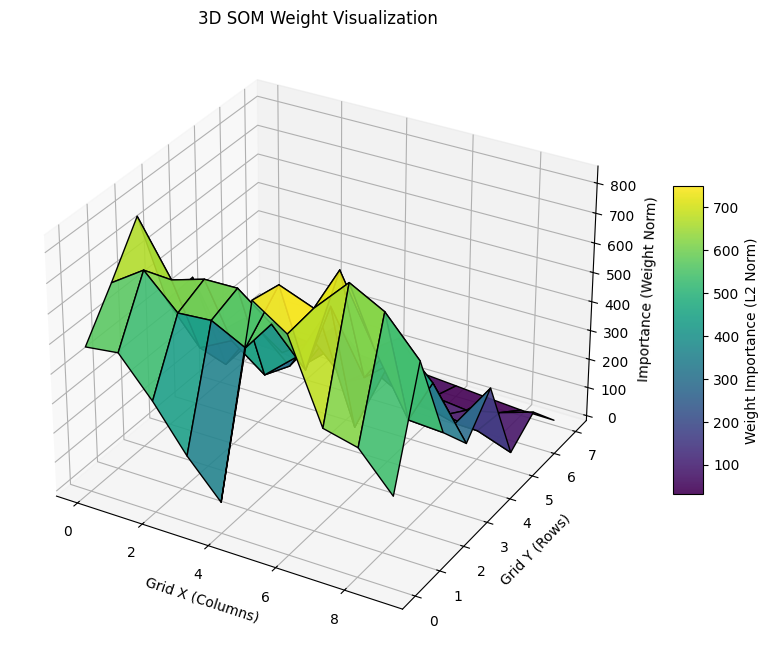
Experimental Results
We evaluated NPSOM on two major multiomics datasets: TCGA-BRCA (Breast Invasive Carcinoma) and ROSMAP (Alzheimer's detection).
Comparison with State-of-the-Art
| Method | BRCA Acc. | ROSMAP Acc. |
|---|---|---|
| KNN | 0.783 | 0.651 |
| RF | 0.768 | 0.754 |
| LASSO | 0.772 | 0.755 |
| XGBoost | 0.791 | 0.764 |
| MOGONET | 0.806 | 0.800 |
| SEGCN | 0.840 | 0.792 |
| MOSEGCN | 0.867 | 0.830 |
| NPSOM+Proto Net (Ours) | 0.8707 ± 0.012 | 0.8345 ± 0.016 |
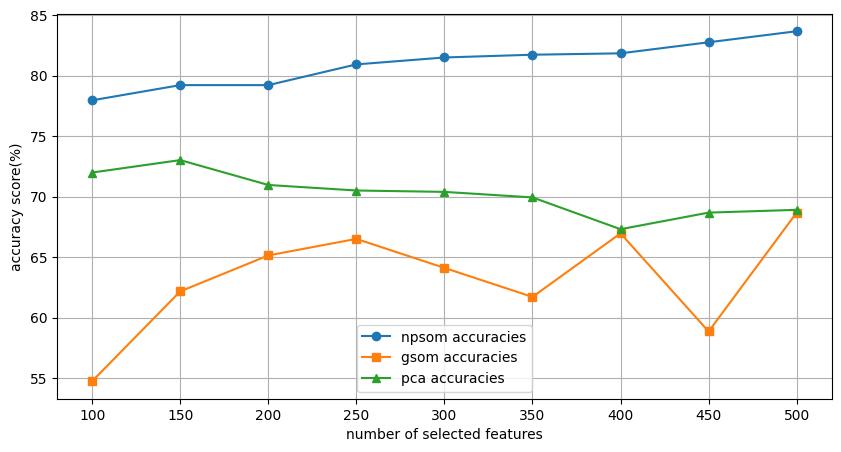
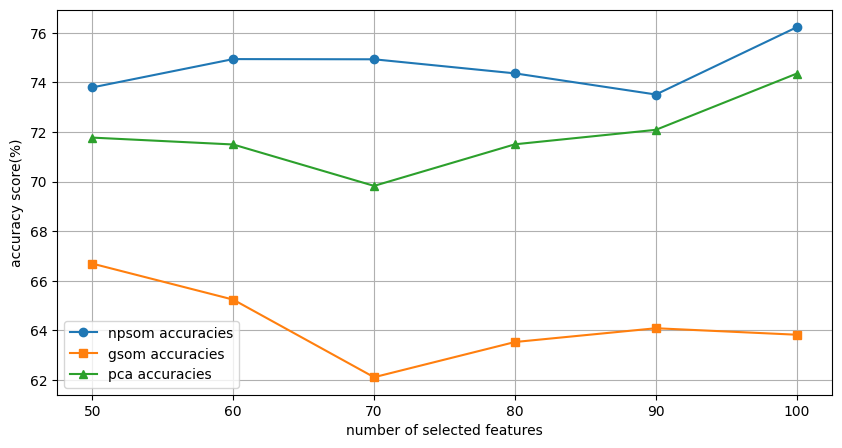
Ablation Study
To understand the contribution of each neuroplasticity component, we conducted an ablation study.
| Model | BRCA Acc (%) | ROSMAP Acc (%) |
|---|---|---|
| Full Model | 87.07 ± 1.2 | 83.45 ± 1.6 |
| Without Neurogenesis | 68.74 ± 4.3 | 72.07 ± 2.7 |
| Without Forgetting | 82.25 ± 5.2 | 74.32 ± 5.7 |
| Without Synaptic Decay | 83.62 ± 3.1 | 72.09 ± 4.4 |
Conclusion
NPSOM demonstrates that incorporating neuroplasticity features into unsupervised learning models significantly enhances their ability to handle complex, high-dimensional multiomics data. By dynamically adjusting to data distributions, NPSOM provides a more robust and biologically plausible framework for feature selection and data analysis in bioinformatics.